One of the big challenges preventing routine application of comprehensive 2D-LC (LC×LC) arises from the complex data structure produced by each experiment. Indeed, with the first-dimension separation only scarcely sampled by the modulator, only a handful of datapoints are available to describe the information [1]. In contrast, second-dimension signals generally offer a surplus of datapoints. As a result, describing two-dimensional peaks – in particular those which co-elute with other species, can be rather difficult.
This becomes a significant issue when the chromatograms comprise a vast number of eluting analytes, and an actual bottleneck when chromatograms of a large number of samples must be compared.
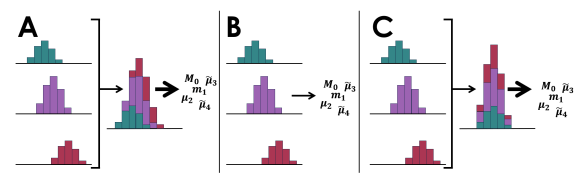
Figure 1. Assessing peaks in LC×LC also becomes a problem during computational assessment. Due to the fact that the modulator samples each first-dimension peak multiple times, the peak is divided over different second-dimension modulations. As modern 2D-LC methods often utilize advanced composition programs (e.g. shifting gradients), the peak will elute differently for each cut. With furthermore the risk of undersampling, Molenaar et al. investigated the three strategies to evaluate peak statistical moments. Reproduced with permission from [2].
This problem is even more critical for analyte mixtures where species do not resolve as individual peaks but rather as broad envelopes representing distributions of co-eluting peaks (i.e. polymer separations).
Active for the UNMATCHED project, Stef Molenaar works to develop algorithms to improve data analysis and method development of 2D-LC polymer separations. To cross the bridge to polymer data, Molenaar first targeted the already challenging case of resolved “small molecules”.
Meanwhile, the group of Dwight Stoll (Gustavus Adolphus College, MN, United States) specializes in the development of highly-optimized state-of-the-art 2D-LC bioseparations [3].

Figure 2. The algorithm developed by Molenaar et al. was able to automatically track and analyze the majority of the peaks in different LC×LC-MS measurements of monoclonal-antibody peptides. Reproduced with permission from [2].
We therefore teamed up with the group of Dwight Stoll and developed a peak-tracking algorithm to evaluate LC×LC-MS datasets and combine their information [2]. The algorithm computes statistical moments for each dimension and consults mass spectra in order to build an assessment. To reduce the number of evaluations and save computational resources, the algorithm compares the retention patterns between the supplied chromatogram. The work was based on an earlier work by Molenaar [4].
The latter is particularly important because this renders the algorithm useful for method-optimization programs where different methods are used to record chromatograms of the same sample in order to allow retention modelling.
While noting that the performance of the algorithm heavily relied on peak-detection algorithms to locate the peaks, the majority of the sample components were well tracked.
These and other conclusions are detailed in the article which was published in Journal of Chromatography A as open-access article. This means you can download and read it for free. Meanwhile, we decided to continue this project, so we hope to be able to report more about this at a later stage.
References
[1] Recent Developments in Two-Dimensional Liquid Chromatography: Fundamental Improvements for Practical Applications
B.W.J. Pirok, D.R. Stoll and P.J. Schoenmakers, Anal. Chem., 2019, 91(1), 240-263, DOI: 10.1021/acs.analchem.8b04841 OPEN ACCESS
[2] Peak-tracking algorithm for use in comprehensive two-dimensional liquid chromatography – application to monoclonal-antibody peptides, S.R.A. Molenaar, T.A. Dahlseid, G. Leme, D.R. Stoll, P.J. Schoenmakers, B.W.J. Pirok, J. Chromatogr. A, 2021, 1639, 461922, DOI: 10.1016/j.chroma.2021.461922 OPEN ACCESS
[3] High resolution two-dimensional liquid chromatography coupled with mass spectrometry for robust and sensitive characterization of therapeutic antibodies at the peptide level, D.R. Stoll, H.R. Lhotka, D.C. Harmes, B. Madigan, J.J. Hsiao, G. O. Staples, J Chromatogr. B, 2019, 1134–1135, 121832, DOI: 10.1016/j.chroma.2021.461922.
[4] Peak-Tracking Algorithm for Use in Automated Interpretive Method-Development Tools in Liquid Chromatography
B.W.J. Pirok, S.R.A. Molenaar, L.S. Roca and P.J. Schoenmakers, Anal. Chem., 2018, 90(23), 14011-14019, DOI: 10.1021/acs.analchem.8b03929 OPEN ACCESS

The Author
Stef Molenaar conducted his PhD in Amsterdam and his research was specifically aimed towards the development of computational strategies for polymer LC×LC data analysis and method optimization. He defended his PhD in 2023.

